BIGNON Emmanuelle;A QIU Yuya;A SMIRNOVA Ekaterina;B SCHULTZ Patrick;B PAPAI Gabor;B BEN SHEM AdamB
A) Université de Lorraine and CNRS, UMR 7019 LPCT, Nancy, France; B) Department of Integrated Structural Biology, IGBMC, CNRS and INSERM, UMR 7104-UMR-S 1258, Illkirch, France
emmanuelle.bignon@univ-lorraine.fr
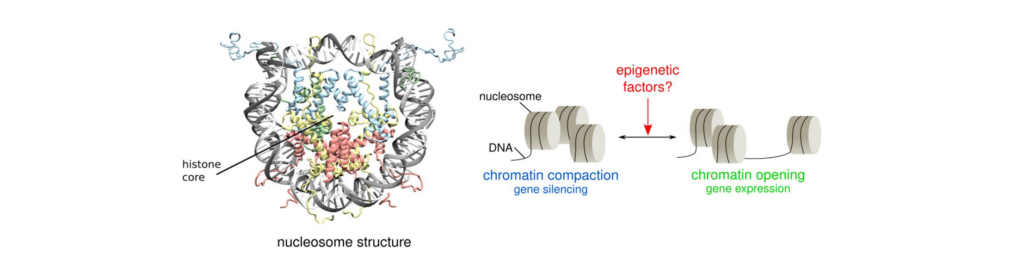
Gene activity is tightly controlled by reversible chemical modifications called epigenetic marks, which are of various types and modulate gene accessibility without affecting the DNA sequence. Major advances come from investigations of such structural regulation at the first level of compaction of DNA, the so-called nucleosome, that is composed of ~146 base pairs of DNA wrapped around an octamer of histone proteins [1]. Indeed, post-translational modifications (PTMs) of histone proteins play are major epigenetic mechanisms. For example, lysine methylation usually promotes gene silencing while acetylation are marks of gene expression. Understanding the molecular mechanisms driving such epigenetic regulation is an active field of research, and many aspects remain to be described, such as how dedicated enzymes regulate the formation and removal of histone PTMs, and how these PTMs impact the nucleosome architecture through specific structural signatures. Sirtuin 6 (SIRT6) is an NAD+-dependent histone H3 deacetylase that is prominently found associated with chromatin, attenuates transcriptionally active promoters and regulates DNA repair, metabolic homeostasis and lifespan. Unlike other sirtuins, it has low affinity to free histone tails but demonstrates strong binding to nucleosomes. It is poorly understood how SIRT6 docking on nucleosomes stimulates its histone deacetylation activity. Combining extended MD simulations to cryogenic electron microscopy (cryoEM) approaches, we revealed the structure and dynamics of the Sirtuin 6 (SIRT6) de-acetylase complexed to a nucleosome core particle [2,3]. We show that in this mode of interaction, the active site of SIRT6 is perfectly poised to catalyze deacetylation of the H3 histone tail and that the partial unwrapping of the DNA allows even lysines close to the H3 core to reach the enzyme, providing new insights into the potential dynamics of SIRT6 bound to a nucleosome, that help explain its substrate specificity.
Keywords: DNA compaction; epigenetics; histones post-translational modifications; MD simulations.
References:
- Cutter, A. R., & Hayes, J. J. (2015). A brief review of nucleosome structure. FEBS letters, 589(20), 2914-2922.
- Smirnova, E., Bignon, E., Schultz, P., Papai, G., & Shem, A. B. (2024). Binding to nucleosome poises human SIRT6 for histone H3 deacetylation. Elife, 12, RP87989.
- Qiu, Y., Papai, G., Ben Shem, A., Bignon, E. (2025). Specific binding modes of SIRT6 C-terminal domain to the nucleosome core particle influence DNA unwrapping and lysine accessibility. BioRxiv.


